<p>Construction of the front loader [own work]: 1—attachment bracket to the tractor; 2—parallel guide long; 3—frame; 4—actuator responsible for lifting the whole structure; 5—actuator responsible for moving the frame; 6—triangle connecting the frame, actuator, and parallel guide long; 7—parallel guide short; 8—hanger; and 9—frame for attaching the attachment.</p> Full article ">Figure 2
<p>View of a 3D model of an extending front loader [own work].</p> Full article ">Figure 3
<p>Force distribution in bottom position without and with extension [own work].</p> Full article ">Figure 4
<p>Von Misses stress distribution of the loader arm without extension (number of mesh elements 49,740, number of nodes 101,251) [own work].</p> Full article ">Figure 5
<p>Detailed view of the most stressed areas of the loader arm without extension [own work].</p> Full article ">Figure 6
<p>Deflection of loader arm structure without extension [own work].</p> Full article ">Figure 7
<p>Von Misses stress distribution of extending loader arm (number of mesh elements 52,397, number of nodes 107,068) [own work].</p> Full article ">Figure 8
<p>Detailed view of the most stressed areas of extending loader arm [own work].</p> Full article ">Figure 9
<p>Deflection of the extending loader arm structure [own work].</p> Full article ">Figure 10
<p>EURO frame deflection analysis [own work].</p> Full article ">Figure 11
<p>Deflection analysis of the frame link, long straight rod, and work tool control actuator (number of elements 26,214, number of nodes 47,632) [own work].</p> Full article ">Figure 12
<p>Deflection analysis of a parallel guide short (number of elements 110,630, number of nodes 182,340) [own work].</p> Full article ">Figure 13
<p>Hanger deflection analysis (number of elements 52,828, number of nodes 87,368) [own work].</p> Full article ">Figure 14
<p>Conceptual model of extension control using a hydraulic motor and helical gearbox [own work].</p> Full article ">Figure 15
<p>View of the mounted loader arm extension drive system [own work].</p> Full article ">
<p>Processing flow chart of aquatic products.</p> Full article ">Figure 2
<p>Schematic diagram of infrared drying.</p> Full article ">Figure 3
<p>Application of computer vision technology in drying process.</p> Full article ">Figure 4
<p>Schematic diagram of the HSI system used in the drying process.</p> Full article ">
<p>Block diagram of VLL.</p> Full article ">Figure 2
<p>Block diagram of VLL with serially assisted PLL.</p> Full article ">Figure 3
<p>Block diagram of VLL with parallelly assisted PLL.</p> Full article ">Figure 4
<p>Structure of second-order VLL loop filter.</p> Full article ">Figure 5
<p>Structure diagram of HD-VLL.</p> Full article ">Figure 6
<p>Structure of multi-carrier NCO.</p> Full article ">Figure 7
<p>Structure of HD-VLL-NCO.</p> Full article ">Figure 8
<p>Implementation for HD-VLL-NCO.</p> Full article ">Figure 9
<p>The tracking results of different bandwidths for the HD-VLL. (<b>a</b>) The velocity estimate curves of different bandwidths for the HD-VLL. (<b>b</b>) The velocity error curves of different bandwidths for the HD-VLL.</p> Full article ">Figure 10
<p>The tracking results of the VLL, the VLL in [<a href="#B11-electronics-13-01794" class="html-bibr">11</a>], and the HD-VLL with a jerk of 10 g/s. (<b>a</b>) The velocity estimate curve. (<b>b</b>) The velocity error curves.</p> Full article ">Figure 11
<p>The tracking results of the VLL, the VLL in [<a href="#B11-electronics-13-01794" class="html-bibr">11</a>], and the HD-VLL with a velocity of [20<span class="html-italic">sin</span>(10<span class="html-italic">t</span>) + 30]. (<b>a</b>) The velocity estimate curve. (<b>b</b>) The velocity error curves.</p> Full article ">Figure 12
<p>The tracking results of the VLL, the VLL in [<a href="#B11-electronics-13-01794" class="html-bibr">11</a>], and the HD-VLL with a velocity of [20<span class="html-italic">sin</span>(2<span class="html-italic">t</span><sup>2</sup>) + 30]. (<b>a</b>) The velocity estimate curve. (<b>b</b>) The velocity error curve.</p> Full article ">Figure 13
<p>The tracking results of different bandwidths for the VLL. (<b>a</b>) The velocity estimate curves of different bandwidths for the VLL. (<b>b</b>) The velocity error curves of different bandwidths for the VLL.</p> Full article ">Figure 14
<p>The tracking results of the HD-VLL and VLL in the sinusoidal motion case. (<b>a</b>) The velocity estimate curves of the HD-VLL and VLL in the sinusoidal motion case. (<b>b</b>) The velocity error curves of the HD-VLL and VLL in the sinusoidal motion case.</p> Full article ">Figure 15
<p>The tracking results of the HD-VLL and VLL in the fixed jerk case. (<b>a</b>) The velocity estimate curves of the HD-VLL and VLL in the fixed jerk case. (<b>b</b>) The velocity error curves of the HD-VLL and VLL in the fixed jerk case.</p> Full article ">Figure 16
<p><b>The</b> PLL tracking results of the HD-VLL-SA-PLL and VLL-SA-PLL. (<b>a</b>) The PLL discrimination curves. (<b>b</b>) The PLL filtering curves.</p> Full article ">Figure 17
<p>The tracking results of the HD-VLL-NCO and HD-VLL (5 Hz). (<b>a</b>) The velocity estimate curves. (<b>b</b>) The velocity error curves.</p> Full article ">
<p>Structure of 6,6′-dihydroxythiobinupharidine (DTBN).</p> Full article ">Figure 2
<p>Survival of DTBN-treated <span class="html-italic">Lm</span> promastigotes. First, 1 × 10<sup>6</sup>/well <span class="html-italic">Leishmania major</span> promastigotes in 1 ml were added to 96-well plates in a 200 µL final volume (200,000 parasites/well). Each well was treated with paromomycin (Paro.) as the gold standard, DTBN 0.1 µg/mL or DTBN 0.2 µg/mL or vehicle. Then, 48 h later, the motile (live) promastigotes were counted out of 100 parasites and the percentage of viable parasites was confirmed on a hemocytometer with trypan blue. The surviving promastigotes are presented as a percentage of untreated live promastigotes. Mean and SD were calculated.</p> Full article ">Figure 3
<p>Transmission (TEM) and scanning electron microscopy (SEM) images of <span class="html-italic">L. major</span> promastigotes treated with DTBN. <span class="html-italic">L. major</span> promastigotes were treated with growth media or 0.1 µg/mL DTBN for 24 h. The cells were then washed with PBS and processed as described for EM evaluation. (<b>A1</b>) Control, untreated promastigotes with SEM pictures (bar 5 mm), (<b>A2</b>) Untreated promastigotes, with TEM pictures. (<b>B1</b>) SEM pictures of treated promastigotes at two magnifications (<b>left B1</b>) (5.55K (bar 15 mm) and (<b>right B1</b>) 7.5K (bar 7.5 mm) magnification (bar 7.5 mm)). Arrows show damaged promastigotes, some lack flagella or show cell body and organelle damage. (<b>B2</b>) TEM of promastigotes treated with 0.1 µg/mL DTBN. M—mitochondria; N—nucleus; F—flagella; K—kinetoplast.</p> Full article ">Figure 4
<p>DTBN treatment of infected macrophages reduces the amount of intracellular <span class="html-italic">L. major</span> amastigotes. First, 24-well plates were covered with slides. Then, 3 × 10<sup>5</sup>/well C3H peritoneal macrophages were left to adhere for 3 h on glass-slide-covered wells. Next, 1 × 10<sup>6</sup>/well <span class="html-italic">L. major</span> promastigotes were added to each well (in triplicate). After 24 h, cells were treated (<b>A</b>) vehicle, (<b>B</b>) with paromomycin (Paro) 200 µg/mL, (<b>C</b>) DTBN 2 µg/mL or (<b>D</b>) DTBN 5 µg/mL. After 72 h, the slides were Giemsa stained and the intracellular amastigotes are shown by the arrow. Pictures were taken at a ×1000 magnification. (<b>E</b>) In parallel, 72 h later, equally treated macrophage cultures were collected, and cDNA was synthesized. The DNA samples were run in multiplex hydrolysis probe-based real-time PCR (mqRT-PCR). <span class="html-italic">L. major</span> cDNA relative amounts as compared to the positive control paromomycine were calculated based on a promastigote DNA calibration curve. The results were normalized to vehicle-treated values. The <span class="html-italic">Leishmania</span>-infected macrophage cDNA sample value was 1. Thus, 2 µg/mL was not effective and 3 and 5 µg/mL showed a dose response of about 10% and 70% effectiveness.</p> Full article ">Figure 5
<p><span class="html-italic">Nuphar lutea</span> semi-purified extract (NUP.E) reduces the size of <span class="html-italic">Lm</span> wounds in treated mice. First, 1 × 10<sup>8</sup>/100 µL <span class="html-italic">L. major</span> promastigotes were injected into the tail base of 40 Balb/c male mice. Ten days later, following the appearance of the wound, 20 mg/kg/mouse NUP.E or acidic water (vehicle) was IP injected daily into mice for 8 days. Small wounds were defined as <0.1 cm<sup>2</sup>; large wounds as >0.1 cm<sup>2</sup>. The wound size was determined by the Digimizer program. Pictures of the wounds were taken at days 0 and 8 after treatment as well as 5 and 30 days post-treatment. (<b>A</b>) Representative pictures of differences in wound size between treated and control groups. (<b>B</b>) Proportion of mice with small/large wound area after 8 days of treatment. (<b>C</b>) Proportion of mice with small/large wound area 5 days post-treatment. (<b>D</b>) Wound area scattering of treated mice (each dot represents the wound area of one mouse in the experiment) as compared to control, 5 days post-treatment (<span class="html-italic">t</span>-test; <span class="html-italic">p</span> < 0.0016, ** <span class="html-italic">p</span> < 0.01). (<b>E</b>) Wound area scattering of treated mice as compared to control, 30 days post-treatment. No significant difference in wound size was observed between the two groups. Mice were sacrificed when the wound area reached 1.5 cm<sup>2</sup>.</p> Full article ">Figure 6
<p>Reduction in in vivo wound size of <span class="html-italic">L. major</span>-infected mice by intra-lesion (IL) injection of DTBN or vehicle. First, 1 × 10<sup>8</sup>/100 µL <span class="html-italic">L. major</span> promastigotes were injected into the mouse tail base. Next, 10 days afterward, the lesions appeared and 20 µg/40 µL of DTBN or vehicle (diluted DMSO) (n = 18) was IL injected. As a positive control, 20 mg/kg/mouse Pentostam (PENT) (n = 18) was IP injected. All treatments were given once a day for 15 days (R1) and the treatment was then stopped for 12 days and resumed for 7 more days (R2). Wound pictures were taken 30 days post-R2. (<b>A</b>) Increasing wound size from the smallest to the largest in each group, 30 days post-R2. (<b>B</b>) Wound size 30 days post-R2 of mice treated with DTBN or Pentostam as compared to the vehicle group. (Mean and SEM were calculated by Anova test; <span class="html-italic">p</span> < 0.0128, * <span class="html-italic">p</span> < 0.05). (<b>C</b>) Wound size distribution 30 days post-R2. (<b>D</b>) Proportion of mice with large/small wounds 30 days post-R2. The number of mice in each group was 18 (small wound: cm<sup>2</sup> < 0.1; big wound: cm<sup>2</sup> > 0.1 The wound size was analyzed by the Digimizer program). (<b>E</b>) The proportion of mice who survived in each group (18 mice/group) was determined at the endpoint of the experiment, 48 days post-R2. Mice were sacrificed when the wound area reached 1.5 cm<sup>2</sup>. (<b>F</b>) Wound size was compared at 48 days post-R2 to the wound size at 30 days post-R2. Unchanged wound size or size reduction was considered a cured wound.</p> Full article ">
<p><span class="html-italic">Oscamta3</span> exhibited increased resistance to <span class="html-italic">M. oryzae</span>. (<b>a</b>) Fifteen-day-old ZH11 and <span class="html-italic">oscamta3</span> seedlings inoculated with <span class="html-italic">M. oryzae</span> by spraying. Images were taken at 5 days post inoculation (dpi). Bar = 1 cm. (<b>b</b>) Fungal biomass of spraying-inoculated leaves was measured to quantify relative fungal growth in ZH11 and <span class="html-italic">oscamta3</span>. Data are presented as the means ± SEs (<span class="html-italic">n</span> = 3). Lowercase letters indicate statistically significant differences (<span class="html-italic">p</span> < 0.05; one-way ANOVA). (<b>c</b>) Twenty-five-day-old ZH11 and <span class="html-italic">oscamta3</span> seedlings inoculated with <span class="html-italic">M. oryzae</span> by punch inoculation. Images were taken at 7 dpi. Bar = 1 cm. (<b>d</b>) Fungal biomass of punch-inoculated leaves was measured to quantify relative fungal growth in ZH11 and <span class="html-italic">oscamta3</span>. Data are presented as the means ± SEs (<span class="html-italic">n</span> = 3). Lowercase letters indicate statistically significant differences (<span class="html-italic">p</span> < 0.05; one-way ANOVA). (<b>e</b>) <span class="html-italic">Oscamta3</span> showed obvious lesion-mimic cell death at the heading stage. Bar = 1 cm.</p> Full article ">Figure 2
<p><span class="html-italic">Oscamtapl</span> exhibited increased resistance to <span class="html-italic">M. oryzae</span>. (<b>a</b>) Fifteen-day-old ZH11 and <span class="html-italic">oscamtapl</span> seedlings inoculated with <span class="html-italic">M. oryzae</span> Zhong1 by spraying. Images were taken at 5 dpi. Bar = 1 cm. (<b>b</b>) Fungal biomass of spraying-inoculated leaves was measured to quantify relative fungal growth in ZH11 and <span class="html-italic">oscamtapl</span>. Data are presented as the means ± SEs (<span class="html-italic">n</span> = 3). Lowercase letters indicate statistically significant differences (<span class="html-italic">p</span> < 0.05; one-way ANOVA). (<b>c</b>) Twenty-five-day-old ZH11 and <span class="html-italic">oscamtapl</span> seedlings inoculated with <span class="html-italic">M. oryzae</span> Zhong1 by punch inoculation. Images were taken at 7 dpi. Bar = 1 cm. (<b>d</b>) Fungal biomass of punch-inoculated leaves was measured to quantify relative fungal growth in ZH11 and <span class="html-italic">oscamtapl</span>. Data are presented as the means ± SEs (<span class="html-italic">n</span> = 3). Lowercase letters indicate statistically significant differences (<span class="html-italic">p</span> < 0.05; one-way ANOVA). (<b>e</b>) <span class="html-italic">Oscamtapl</span> did not show an obvious hypersensitive response at the heading stage. Bar = 1 cm.</p> Full article ">Figure 3
<p><span class="html-italic">Oscamta3/pl</span> exhibited increased resistance to <span class="html-italic">M. oryzae</span>. (<b>a</b>) Fifteen-day-old ZH11, <span class="html-italic">oscamta3</span>, <span class="html-italic">oscamtapl</span>, and <span class="html-italic">oscamta3/pl</span> seedlings inoculated with <span class="html-italic">M. oryzae</span> by spraying. Images were taken at 5 dpi. Bar = 1 cm. (<b>b</b>) Fungal biomass of spraying-inoculated leaves was measured to quantify relative fungal growth in ZH11, <span class="html-italic">oscamta3</span>, <span class="html-italic">oscamtapl</span>, and <span class="html-italic">oscamta3/pl</span>. Data are presented as the means ± SEs (<span class="html-italic">n</span> = 3). Lowercase letters indicate statistically significant differences (<span class="html-italic">p</span> < 0.05; one-way ANOVA). (<b>c</b>) Twenty-five-day-old ZH11, <span class="html-italic">oscamta3</span>, <span class="html-italic">oscamtapl</span>, and <span class="html-italic">oscamta3/pl</span> seedlings inoculated with <span class="html-italic">M. oryzae</span> by punch inoculation. Images were taken at 7 dpi. (<b>d</b>) Fungal biomass of punch-inoculated leaves was measured to quantify relative fungal growth in ZH11 and <span class="html-italic">oscamta3/pl</span>. Data are presented as the means ± SEs (<span class="html-italic">n</span> = 3). Lowercase letters indicate statistically significant differences (<span class="html-italic">p</span> < 0.05; one-way ANOVA). (<b>e</b>) <span class="html-italic">Oscamta3/pl</span> plants showed obvious lesion-mimic cell death at the heading stage. Bar = 1 cm.</p> Full article ">Figure 4
<p>OsCAMTAPL interacted with the TIG-ANK domain of OsCAMTA3. (<b>a</b>) The different domains of OsCAMTA3 are shown. OsCAMTA3 contains the CG-1 (the gray area), TIG-ANK(the cyan-yellow area) and IQ (the blue area) domains. The numbers indicate the positions of amino acid sequences. (<b>b</b>) The TIG-ANK domain of OsCAMTA3 interacted with OsCAMTAPL in a Y2H assay. The coding sequence (CDS) of <span class="html-italic">OsCAMTA3<sup>CG−1</sup></span>, <span class="html-italic">OsCAMTA3<sup>TIG-ANK</sup></span>, or <span class="html-italic">OsCAMTA3<sup>IQs</sup></span> was fused to the vector pGADT7 (AD), and the CDS of <span class="html-italic">OsCAMTAPL</span> was fused to pGBKT7 (BD). The empty vectors pGADT7 (AD) and pGBKT7 (BD) were used as negative controls. Yeast cells containing BD-OsCAMTAPL and AD-OsCAMTA3<sup>TIG-ANK</sup> plasmids were grown on SD-Leu-Trp-His-Ade medium. (<b>c</b>) Images of <span class="html-italic">N. benthamiana</span> leaves with the indicated constructs in LUC assays. The coding sequence (CDS) of <span class="html-italic">OsCAMTA3<sup>TIG-ANK</sup></span> was fused to the C-terminal fragment of firefly luciferase (cLUC), and the CDS of <span class="html-italic">OsCAMTAPL</span> was fused to the N-terminal fragment of firefly luciferase (nLUC). Green fluorescence protein (GFP)-nLUC and Myc-nLUC were used as negative controls. The indicated constructs were transiently coexpressed in 4-week-old <span class="html-italic">N. benthamiana</span> leaves, and the bioluminescence images were captured by a CCD camera. (<b>d</b>) Co-IP assay was performed by transiently coexpressing OsCAMTA3<sup>TIG-ANK</sup>-Myc and OsCAMTAPL-Flag in <span class="html-italic">N. benthamiana</span> leaves. Total protein was extracted and subjected to immunoprecipitation by anti-Flag beads. Immunoblotting analysis was performed with anti-Myc and anti-Flag antibodies. OsCAMTAPL-Flag, GFP-Flag, and OsCAMTA3<sup>TIG-ANK</sup>-Myc are indicated by blue, black, and red asterisks. Full-length blots are presented in <a href="#app1-ijms-25-05049" class="html-app">Supplementary Figure S6</a>.</p> Full article ">Figure 5
<p>Morphology of ZH11 and <span class="html-italic">oscamtas</span> plants at the heading stage. (<b>a</b>) Phenotypes of ZH11 and <span class="html-italic">oscamtas</span> plants at the heading stage. Bars = 10 cm. (<b>b</b>,<b>c</b>) Comparison of plant height and tiller number between ZH11 and <span class="html-italic">oscamtas</span> plants. Data are presented as the means ± SE (<span class="html-italic">n</span> = 10). Lowercase letters indicate statistically significant differences (<span class="html-italic">p</span> < 0.05; one-way ANOVA).</p> Full article ">Figure 6
<p>Identification of DEGs in ZH11 and <span class="html-italic">oscamta3-1</span> by RNA-seq. (<b>a</b>) The number of DEGs between ZH11 and <span class="html-italic">oscamta3-1</span> before inoculation with <span class="html-italic">M. oryzae</span>. (<b>b</b>) <span class="html-italic">OsALDH2B1</span>, <span class="html-italic">OsBIERF1</span>, <span class="html-italic">OsHAC1</span>, and <span class="html-italic">OsSAMS1</span> were chosen to verify the RNA-Seq results by qRT-PCR. Data are presented as the means ± SEs (<span class="html-italic">n</span> = 3). (**; <span class="html-italic">p</span> < 0.01; Student’s <span class="html-italic">t</span>-test). (<b>c</b>) The number of DEGs between ZH11 and <span class="html-italic">oscamta3-1</span> after inoculation with <span class="html-italic">M. oryzae</span>. (<b>d</b>) Venn diagrams showing the number of EDGs before or after inoculation with <span class="html-italic">M. oryzae</span>.</p> Full article ">Figure 7
<p>Identification of the function of DEGs in ZH11 and <span class="html-italic">oscamta3-1</span>. (<b>a</b>) Functional classification of consensus sequence term enrichment analysis of all DEGs before inoculation with <span class="html-italic">M. oryzae</span>. (<b>b</b>) Functional classification of consensus sequence term enrichment analysis of all DEGs after inoculation with <span class="html-italic">M. oryzae</span>.</p> Full article ">Figure 8
<p>OsCAMTA3 activated the expression of OsALDH2B1 by binding to its CGCG motif. (<b>a</b>) OsCAMTA3<sup>CG−1</sup> bound to the DNA fragment of the <span class="html-italic">OsALDH2B1</span> promoter containing the CGCG motif in an EMSA. Sequences of the probe and mutated probe are presented at the top. The wild-type CGCG motif and the mutant AAAA motif have been indicated by blue and red, respectively. Full-length blots are presented in <a href="#app1-ijms-25-05049" class="html-app">Supplementary Figure S9</a>. (<b>b</b>) qRT-PCR analysis of <span class="html-italic">OsALDH2B1</span> expression in ZH11 and <span class="html-italic">oscamta3-1</span> before and after inoculation with <span class="html-italic">M. oryzae</span>. Data are presented as the means ± SEs (<span class="html-italic">n</span> = 3). (**; <span class="html-italic">p</span> < 0.01; Student’s <span class="html-italic">t</span>-test). (<b>c</b>) The transcripts of <span class="html-italic">OsALDH2B1</span> in ZH11, <span class="html-italic">oscamta1/2-1</span>, <span class="html-italic">oscamta3-1</span>, <span class="html-italic">oscamtapl-1</span>, and <span class="html-italic">oscamta3/pl-1</span> mutants. Data are presented as the means ± SEs (<span class="html-italic">n</span> = 3). Lowercase letters represent statistically significant differences (<span class="html-italic">p</span> < 0.05; one-way ANOVA).</p> Full article ">
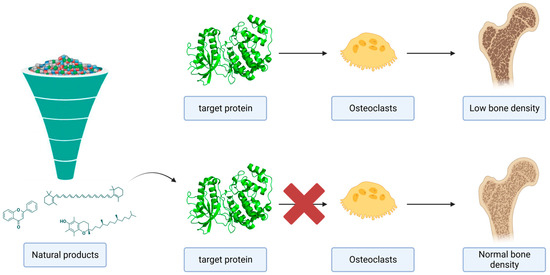
<p>A graphical summary of the role of MAPKs in bone metabolism. The potential inhibition of MAPKs by natural products could suppress the differentiation and activation of the osteoclasts.</p> Full article ">Figure 2
<p>An overview of the workflow implemented for the discovery of natural compounds that have the potential to inhibit MAPKs. The procedure starts with a virtual library enumeration of 180,000 compounds that are filtered based on their reported association with osteoporosis (187 compounds). It then continues with docking calculations, prediction of the interactions of the compounds with the human gut microbiome and assessment of potential clinical trials that have participated and respective EFSA scientific opinions and safety profiles to produce a subset of five ligands in allosteric binding mode. Then, MD simulations are employed on p38b and ERK1 to retain only 2 stably bound molecules, which were subsequently validated in vitro. These compounds undergo testing on additional JNK, p38 and ERK isoforms, followed by a second round of KdELECT assays to identify the natural compound with the potential to modulate the activity of MAPKs. Quercetin emerges as an inhibitor of ERK1, JNK2 and JNK3, suggesting its promise in addressing osteoporosis.</p> Full article ">Figure 3
<p>(<b>A</b>) RMSD of the three replicas of quercetin bound to ERK1 as a function of time; (<b>B</b>) RMSD of the three replicas of the backbone of ERK1 as a function of time; (<b>C</b>) a diagram of the interactions of quercetin with ERK1 (orange), in comparison with the interactions of SCH772984 with ERK1 (blue); (<b>D</b>) the first cluster representative of quercetin bound to ERK1.</p> Full article ">Figure 4
<p>(<b>A</b>) RMSD of the three replicas of naringin bound to p38b as a function of time; (<b>B</b>) RMSD of the three replicas of the backbone of p38b as a function of time; (<b>C</b>) a diagram of the interactions of naringin with p38b (orange), in comparison with the interactions of nilotinib with p38b (blue); (<b>D</b>) the first cluster representative of naringin bound to p38b.</p> Full article ">Figure 5
<p>(<b>A</b>) RMSD of the three replicas of quercetin bound to JNK2 as a function of time; (<b>B</b>) RMSD of the three replicas of the backbone of JNK2 as a function of time; (<b>C</b>) a diagram of the interactions of quercetin with JNK2 (orange), in comparison with the interactions of BIRB-796 with JNK2 (blue); (<b>D</b>) the first cluster representative of quercetin bound to JNK2.</p> Full article ">Figure 6
<p>(<b>A</b>) RMSD of the three replicas of quercetin bound to JNK3 as a function of time; (<b>B</b>) RMSD of the three replicas of the backbone of JNK3 as a function of time; (<b>C</b>) a diagram of the interactions of quercetin with JNK3 (orange), in comparison with the interactions of X3S with JNK3 (blue); (<b>D</b>) the first cluster representative of quercetin bound to JNK3.</p> Full article ">Figure 7
<p>The amount of (<b>A</b>) ERK1, (<b>B</b>) JNK2 and (<b>C</b>) JNK3 measured by qPCR (Signal; <span class="html-italic">y</span>-axis) is plotted against the corresponding quercetin concentration in nM in log10 scale (<span class="html-italic">x</span>-axis).</p> Full article ">Figure 8
<p>Proposed mechanism for quercetin’s inhibition of osteoclastogenesis.</p> Full article ">
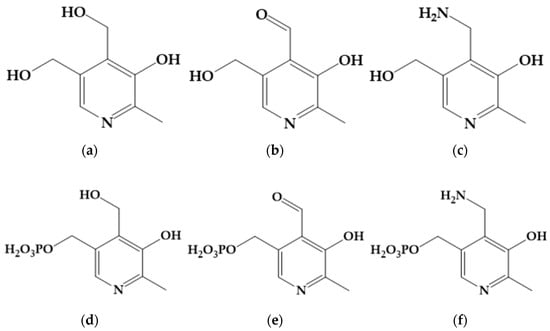
<p>Pyridoxine (<b>a</b>), pyridoxal (<b>b</b>), pyridoxamine (<b>c</b>), pyridoxine 5′-phosphate (<b>d</b>), pyridoxal 5′-phosphate (<b>e</b>), and pyridoxamine 5′-phosphate (<b>f</b>).</p> Full article ">Figure 2
<p>The assumed general structure of the neutral gold(III) complex with the following pyridoxal-derived hydrazones: <b>PL-INH</b> (Ar = 4-pyridinyl); <b>PL-F2H</b> (Ar = 2-furyl); <b>PL-F3H</b> (Ar = 3-furyl); <b>PL-T2H</b> (Ar = 2-thienyl); <b>PL-T3H</b> (Ar = 3-thienyl). Donor atoms involved in complex formation are marked with red, while acceptors of the first and second protons are marked with blue and green, respectively.</p> Full article ">Figure 3
<p>Examples of UV-Vis spectra acquired during the titration of (<b>a</b>) 202.4 µM H[AuCl<sub>4</sub>] + 299.4 µM HClO<sub>4</sub> by 1.657 mM <b>PL-INH</b> + 10.34 mM NaOH; and (<b>b</b>) 200.9 µ H[AuCl<sub>4</sub>] + 298.9 µM HClO<sub>4</sub> by 1.491 mM <b>PL-T2H</b> + 10.84 mM NaOH in water. Initial volume 2.7 mL, 25 to 30 titration points, titrant volume = 10 µL.</p> Full article ">Figure 4
<p>Calculated UV-Vis spectra of individual protonated and complex species of <b>PL-INH</b> (<b>a</b>) and <b>PL-T2H</b> (<b>b</b>). The protonated species spectra are adopted from [<a href="#B47-ijms-25-05046" class="html-bibr">47</a>].</p> Full article ">Figure 5
<p>The <span class="html-italic">C</span><sub>1</sub> symmetry molecular models of neutral ([AuClL]), monoprotonated ([AuCl(HL)]<sup>+</sup> 1, and [AuCl(HL)]<sup>+2</sup>) and <span class="html-italic">bis</span>-protonated ([AuCl(H<sub>2</sub>L)]<sup>+2</sup>) forms of complexes formed with <b>PL-F3H</b> hydrazone. O<sub>p</sub> stands for oxygen of phenyl group in site 3 of pyridoxal moiety; O<sub>c</sub> stands for carbonyl oxygen.</p> Full article ">Figure 6
<p>TD DFT (<b>a</b>,<b>b</b>) and experimental (<b>c</b>,<b>d</b>) UV-Vis spectra of different protonated forms of gold(III) complexes with <b>PLP-F3H</b> (<b>a</b>,<b>c</b>) and <b>PL-F3H</b> (<b>b</b>,<b>d</b>) hydrazones. The red dashed line spectra correspond to single-protonated complexes, where the proton is bound with heterocyclic nitrogen of a <b>PLP</b> ([AuCl(HL)] 1) and <b>PL</b> ([AuCl(HL)]<sup>+</sup> 1) moiety. The theoretical individual bands were described by Lorentz curves with a half-width of 30 nm.</p> Full article ">Figure 7
<p>The HOMO–LUMO diagram for deprotonated [AuClL], monoprotonated [AuCl(HL)]<sup>+</sup> 2, and <span class="html-italic">bis</span>-protonated [AuCl(H<sub>2</sub>L)]<sup>+2</sup> forms of the gold(III) complex with a <b>PL-F3H</b> hydrazone.</p> Full article ">
Open Access Journals
-
IJMS IMPACT
FACTOR
5.6 -
Applied Sciences IMPACT
FACTOR
2.7 -
Sustainability IMPACT
FACTOR
3.9 -
Sensors IMPACT
FACTOR
3.9 -
JCM IMPACT
FACTOR
3.9 -
Materials IMPACT
FACTOR
3.4 -
Molecules IMPACT
FACTOR
4.6 -
Energies IMPACT
FACTOR
3.2 -
Electronics IMPACT
FACTOR
2.9 -
Remote Sensing IMPACT
FACTOR
5.0 -
Cancers IMPACT
FACTOR
5.2 -
Nutrients IMPACT
FACTOR
5.9 -
Mathematics IMPACT
FACTOR
2.4 -
Foods IMPACT
FACTOR
5.2 -
Buildings IMPACT
FACTOR
3.8 -
Polymers IMPACT
FACTOR
5.0 -
Animals IMPACT
FACTOR
3.0 -
Water IMPACT
FACTOR
3.4 -
Plants IMPACT
FACTOR
4.5 -
Agronomy IMPACT
FACTOR
3.7 -
Biomedicines IMPACT
FACTOR
4.7 -
Processes IMPACT
FACTOR
3.5 -
Microorganisms IMPACT
FACTOR
4.5 -
Diagnostics IMPACT
FACTOR
3.6 -
Nanomaterials IMPACT
FACTOR
5.3 -
Viruses IMPACT
FACTOR
4.7 -
Medicina IMPACT
FACTOR
2.6 -
Healthcare IMPACT
FACTOR
2.8 -
Cells IMPACT
FACTOR
6.0 -
Forests IMPACT
FACTOR
2.9 -
Agriculture IMPACT
FACTOR
3.6 -
Land IMPACT
FACTOR
3.9 -
JMSE IMPACT
FACTOR
2.9 -
IJERPH
-
Symmetry IMPACT
FACTOR
2.7 -
Genes IMPACT
FACTOR
3.5 -
Pharmaceutics IMPACT
FACTOR
5.4 -
Coatings IMPACT
FACTOR
3.4 -
Micromachines IMPACT
FACTOR
3.4 -
Pharmaceuticals IMPACT
FACTOR
4.6 -
Atmosphere IMPACT
FACTOR
2.9 -
Children IMPACT
FACTOR
2.4 -
Religions IMPACT
FACTOR
0.8 -
Antioxidants IMPACT
FACTOR
7.0 -
Life IMPACT
FACTOR
3.2 -
Metals IMPACT
FACTOR
2.9 -
Biomolecules IMPACT
FACTOR
5.5 -
Vaccines IMPACT
FACTOR
7.8 -
Education Sciences IMPACT
FACTOR
3.0 -
Minerals IMPACT
FACTOR
2.5 -
Horticulturae IMPACT
FACTOR
3.1 -
Brain Sciences IMPACT
FACTOR
3.3 -
JPM IMPACT
FACTOR
3.4 -
Bioengineering IMPACT
FACTOR
4.6